Labeling carbon backbone amino acid11/18/2023 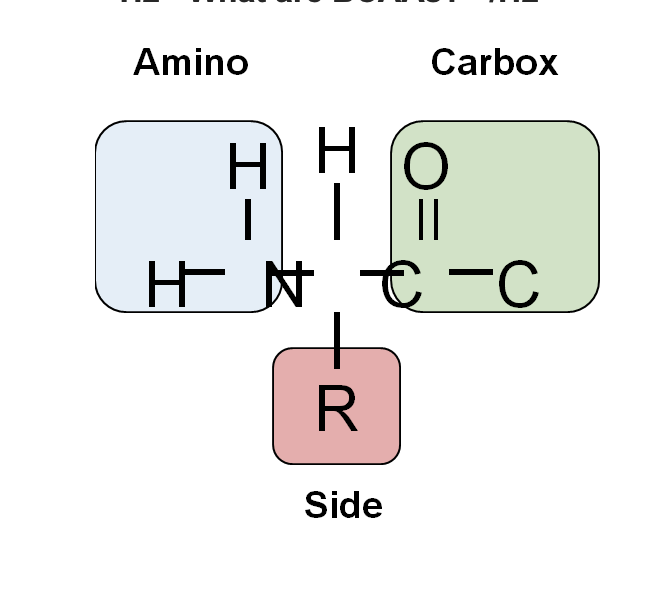
They also contain a side chain that is made up of an R-group (where ‘R’ can denote any alkyl or aryl group). Amino acids are known to contain amine and carboxyl functional groups. 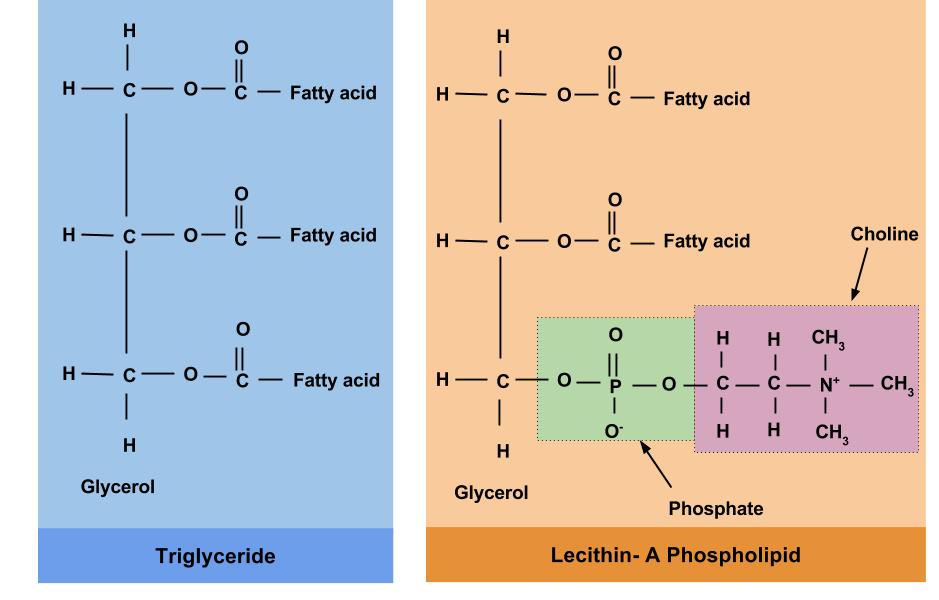 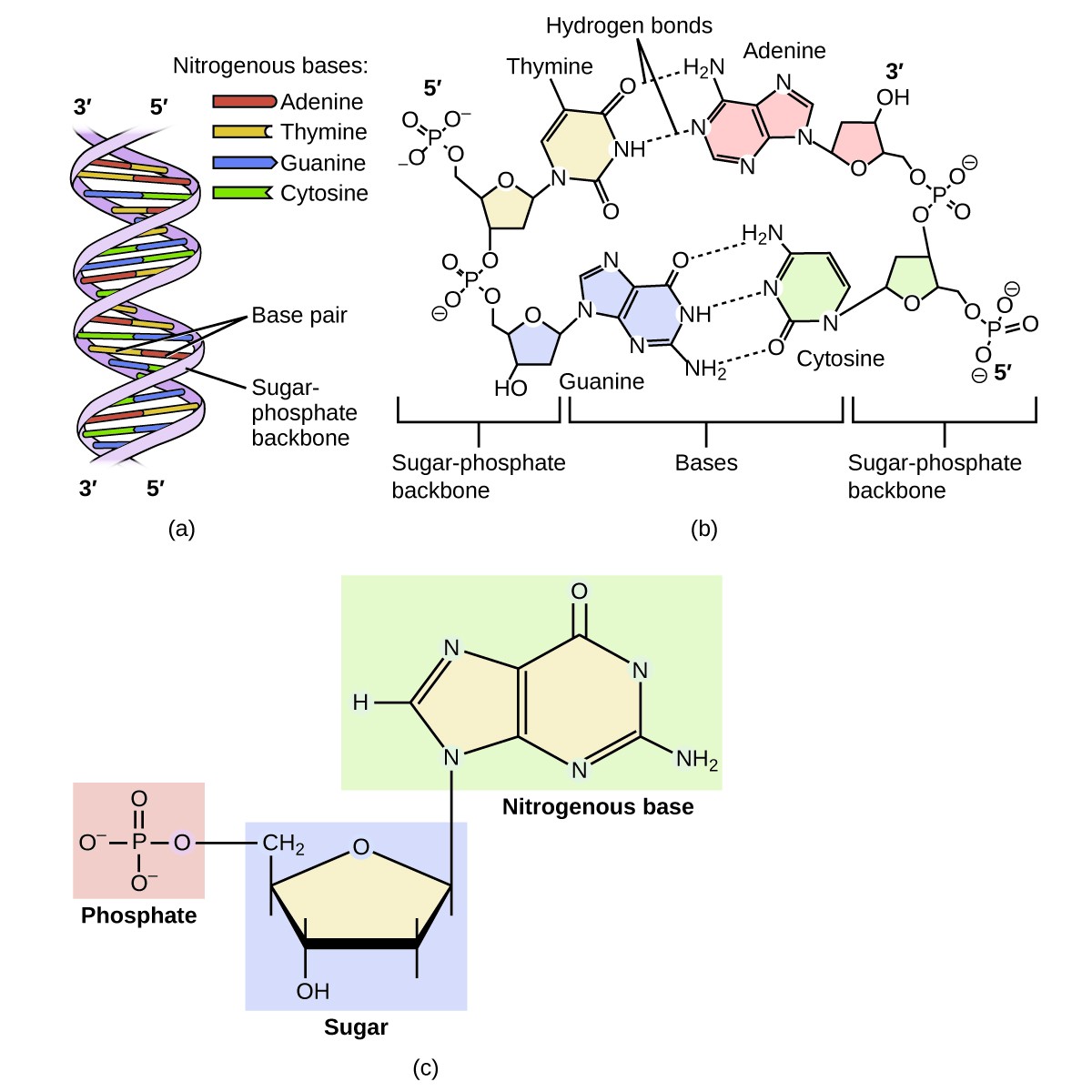
The general formula of an amino acid is R-CH (NH 2 )-COOH. Further studies have since shown that certain electronically stabilized carboxylic acids, such as aryl acetic acids and tertiary carboxylic acids, could undergo reversible decarboxylation–carboxylation in polar aprotic solvents under thermal or photochemical conditions, without the need of transition-metal catalysis, to yield isotopic carbon-enriched carboxylic acids 4, 5, 6, 7. Amino acids are organic compounds that combine to form proteins. In 2019, two independent studies reported the Ni-catalysed decarboxylation–carboxylation of activated esters to yield 13C- and 14C-enriched carboxylic acids 2, 3. In 2018, a method for the Pd-catalysed decarbonylation–carbonylation of acid chlorides was reported, yielding isotopically labelled carboxylic acids 1. Over recent years, several methods have emerged describing the carbon labelling of biologically relevant compounds using the principles of dynamic isotope exchange. These include D-amino acids, homo amino acids, beta-homo amino acids, N-methyl amino acids, alpha-methyl amino acids, non-natural side chain variant amino acids and other unusual amino acids.The preparation of molecular targets, where a native isotope is replaced by a heavier or radiolabelled isotope, is crucial for drug development programmes and medical imaging. TOAC was the first spin label probe incorporated in peptides by means of a peptide bond. 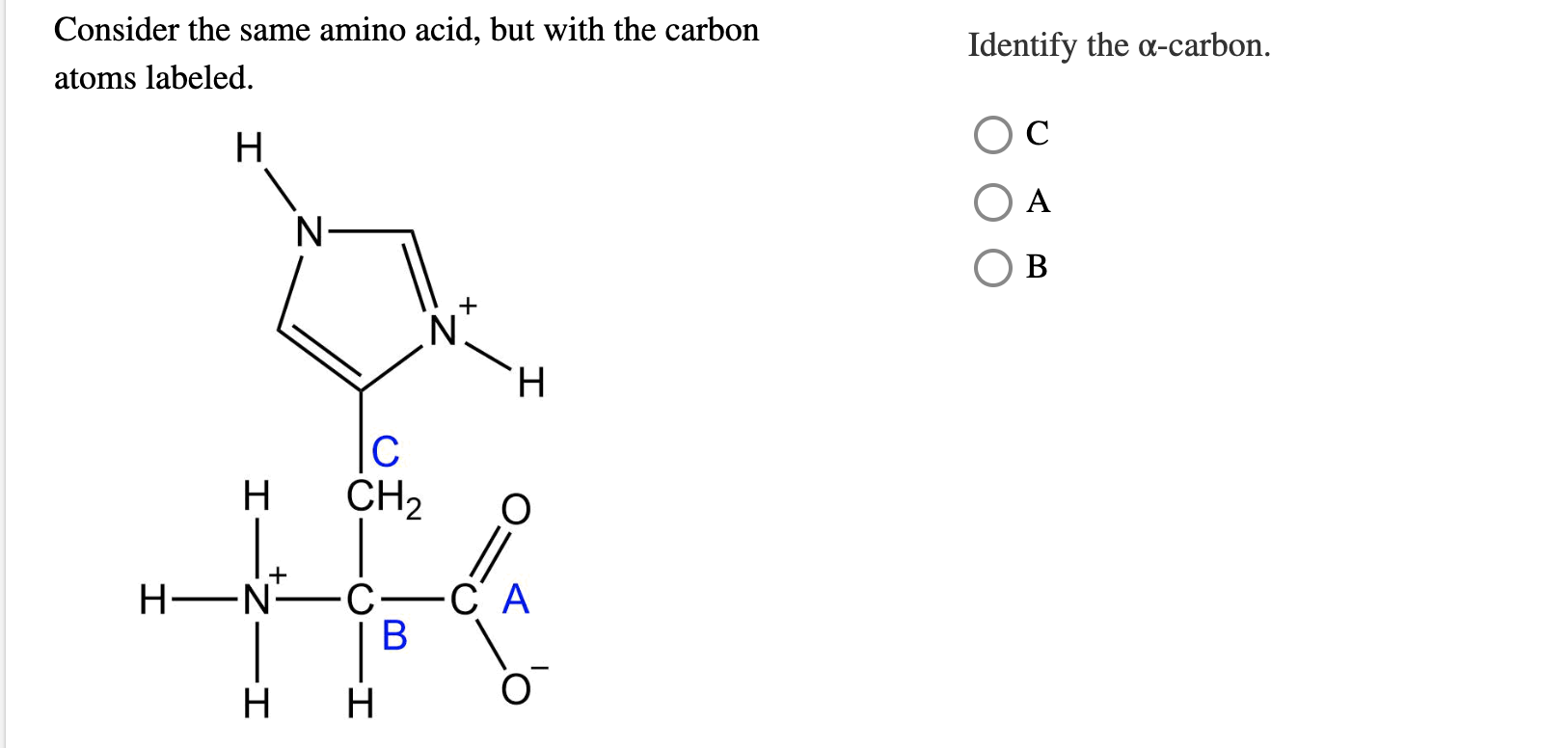
Horizontal lines identify the carbon backbone and side chain res- onances. A peptide bond forms when the carboxylic acid group (R-COOH) of one amino acid reacts with the amine group (R-NH 2) of another.The resulting molecule is an amide with a CN bond (R-C(O)-NH-R). We review work on the paramagnetic amino acid 2,2,6,6-tetramethyl-N-oxyl-4-amino-4-carboxylic acid, TOAC, and its applications in studies of peptides and peptide synthesis. Pepscan offers a large number of different non-natural amino acids, which we routinely incorporate into peptides. All isotope-labeled amino acid derivatives were purchased from Cambridge. Another application is the use of non-natural amino acids for induction or stabilization of secondary structures (α-helices, β-sheets, β-turns). There are many different reasons to incorporate non-natural amino acids, such as for example to enhance affinity, selectivity of stability of peptide drug leads. Metabolic precursors that can be used for isotopic labelling and reverse labelling of the carbon sites in branched-chain aliphatic amino acids. (Be sure to minimize formal charges and draw resonance structures where appropriate.): XeF2O, CIO4-, HNO2, CH3CH(NH2)COOH (amino acid alanine). You can find everything about unusual & non-natural amino acids here.Īpart from the 20 natural L-amino acids, there is a multitude of non-natural or unusual amino acids available that can be built into synthetic peptides. Isotopic labelling using amino acids and metabolites. This page will soon be fully integrated into the Biosynth website. In organic chemistry, this stereochemistry is referred. Label the alpha or central carbon, amino group, carboxyl group, and R group. Pepscan (now Biosynth B.V.) is part of Biosynth's Peptide Division. In the structure illustrated in Figure, the amino group is always to the left side of the alpha carbon. We present a computational method for finding optimal labeling patterns for the backbone assignment of membrane proteins and other large proteins that cannot be assigned by conventional strategies. The monomers of proteins are amino acids.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
